order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Locations
Orders Processing,
Shipping & Receiving,
Warehouse
2 Shaker Rd Suites
B001/B101
Shirley, MA 01464
Production Lab
Floor 6, Suite 620
20700 44th Avenue W
Lynnwood, WA 98036
Telephone Numbers
Tel: +1 (206) 374-1102
Fax: +1 (206) 577-4565
Contact Us
Additional Contact Details
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
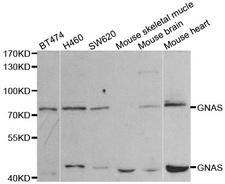
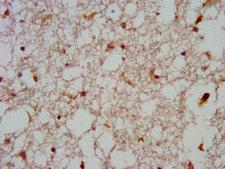
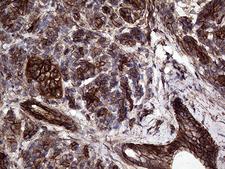
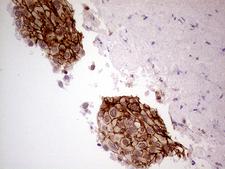
GNAS
GNAS complex locus
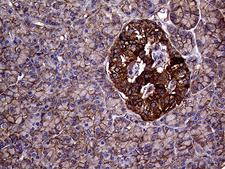
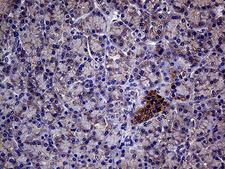
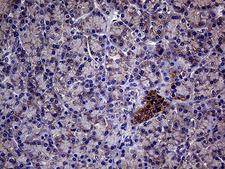
The GNAS1 gene encodes the alpha subunit of the G protein Gs, which couples receptor binding by several hormones to activation of adenylate cyclase (Hayward et al., 1998). This locus has a highly complex imprinted expression pattern. It gives rise to maternally, paternally, and biallelically expressed transcripts that are derived from four alternative promoters and 5' exons. Some transcripts contains a differentially methylated region (DMR) at their 5' exons, and this DMR is commonly found in imprinted genes and correlates with transcript expression. An antisense transcript exists, and this antisense transcript and one of the transcripts are paternally expressed, produce noncoding RNAs, and may regulate imprinting in this region. In addition, one of the transcripts contains a second overlapping ORF, which encodes a structurally unrelated protein - Alex. Alternative splicing of downstream exons is also observed, which results in different forms of the stimulatory G-protein alpha subunit, a key element of the classical signal transduction pathway linking receptor-ligand interactions with the activation of adenylyl cyclase and a variety of cellular reponses. Multiple transcript variants have been found for this gene, but the full-length nature and/or biological validity of some variants have not been determined. Mutations in this gene result in pseudohypoparathyroidism type 1a, pseudohypoparathyroidism type 1b, Albright hereditary osteodystrophy, pseudopseudohypoparathyroidism, McCune-Albright syndrome, progressive osseus heteroplasia, polyostotic fibrous dysplasia of bone, and some pituitary tumors.
| Gene Name: | GNAS complex locus |
| Synonyms: | GNAS, AHO, C20orf45, Extra large alphas protein, GSP, GNAS complex locus, GNASXL, NESP, PHP1B, PHP1C, PHP1A, POH, SCG6, XLalphas, GNAS1, GPSA, GSA, NESP55, Protein ALEX, Secretogranin VI |
| Target Sequences: | NM_080425 NP_536350.2 Q5JWF2 |
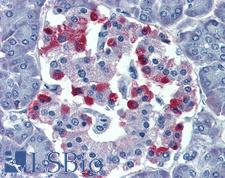
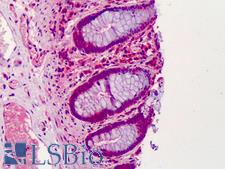
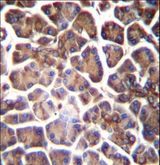
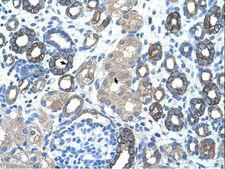
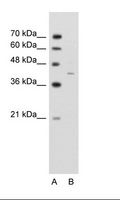
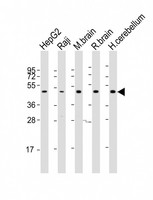
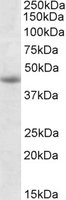
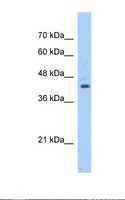

If you do not find the reagent or information you require, please contact Customer.Support@LSBio.com to inquire about additional products in development.










