order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Locations
Orders Processing,
Shipping & Receiving,
Warehouse
2 Shaker Rd Suites
B001/B101
Shirley, MA 01464
Production Lab
Floor 6, Suite 620
20700 44th Avenue W
Lynnwood, WA 98036
Telephone Numbers
Tel: +1 (206) 374-1102
Fax: +1 (206) 577-4565
Contact Us
Additional Contact Details
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Orthomyxoviridae (Influenza)
Influenza viruses are members of the family Orthomyxoviridae, which are enveloped viruses with a segmented negative-sense, single-strand RNA genome. Of the four genera in this family (A, B, C, and D), types A and B are the most clinically relevant because they cause influenza pneumonia with potentially severe complications such as encephalitis, myocarditis, and death. The virus is transmitted by aerosol infection as well as by contaminated surfaces. It enters the body through the nasal and laryngeal mucosa, as well as the lower airways, before spreading systemically to infect a variety of cell types.
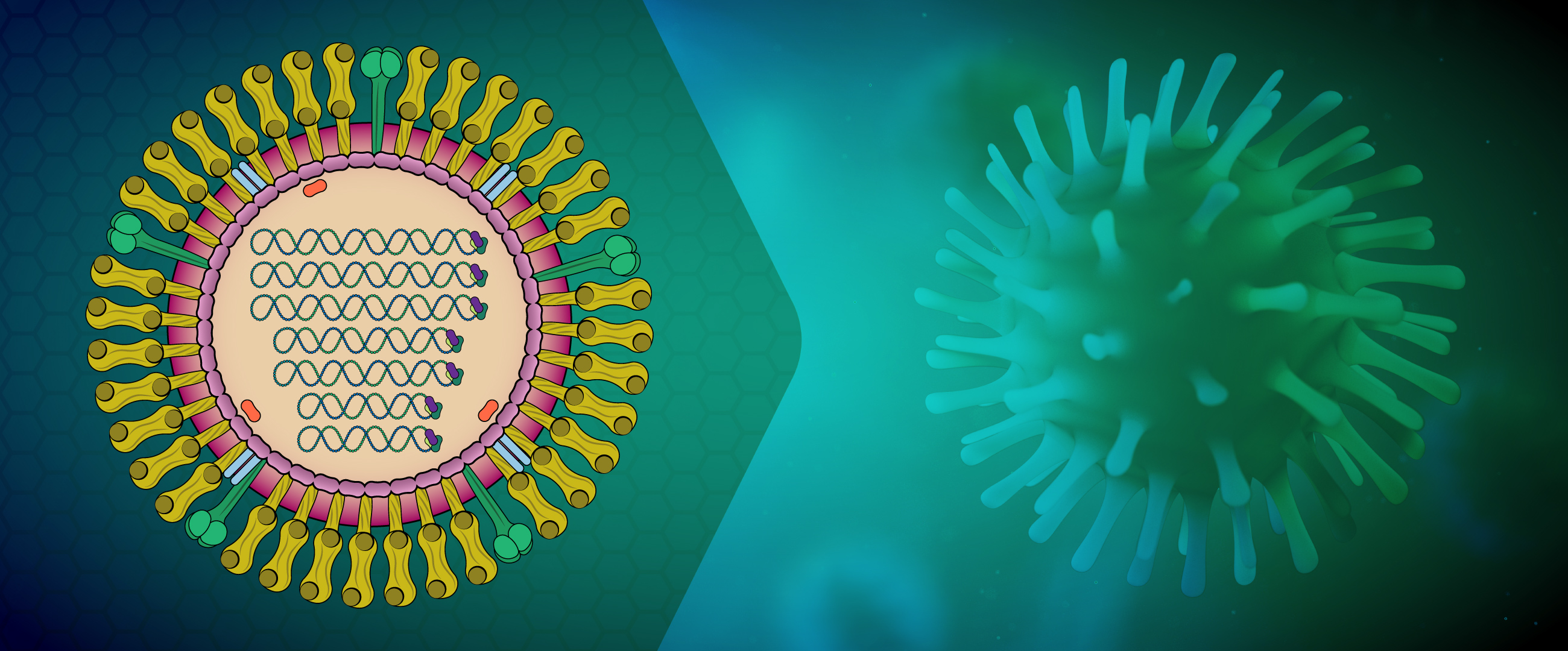
About Orthomyxoviridae
Although Influenza A viruses cause most cases of influenza and are responsible for pandemics, influenza B can cause morbidity during inter-pandemic periods, and up to 23% of influenza cases in a season. It is split into two antigenically distinct phylogenetic lineages (B/Victoria and B/Yamagata). In contrast to influenza A which can be found in multiple species including humans, birds, and pigs, influenza B is typically only found in humans. Treatment for influenza includes antiviral medications such as oseltamivir, marboxil, perimavir, and zanamivir. Vaccination for influenza often protects against strains H1N1 and H3N2 of influenza A and one or two strains of influenza B viruses.
Influenza A virus (IAV) and B virus (IBV) both contain eight viral RNA gene segments, which code for ten essential viral proteins and accessory proteins. IAV and IBV contain the viral lipid membrane glycoproteins HA (hemagglutinin) and NA (neuraminidase) that determine the subtype of influenza virus. HA and NA are both targets for immune responses against the virus, and the NA protein is the target of antiviral drugs such as Relenza and Tamiflu. Also embedded in the lipid membrane is the M2 protein, the target of the antiviral drugs amantadine and rimantadine. Beneath the lipid membrane is M1, the matrix protein that forms the viral capsid, and inside are the viral RNA segments in tight association with polymerase B1 and B2 proteins, PA (polymerase acidic protein), and NP (nucleoprotein). The virion also includes 2 non-structural proteins, NS1 and NEP (nuclear export protein NS2).
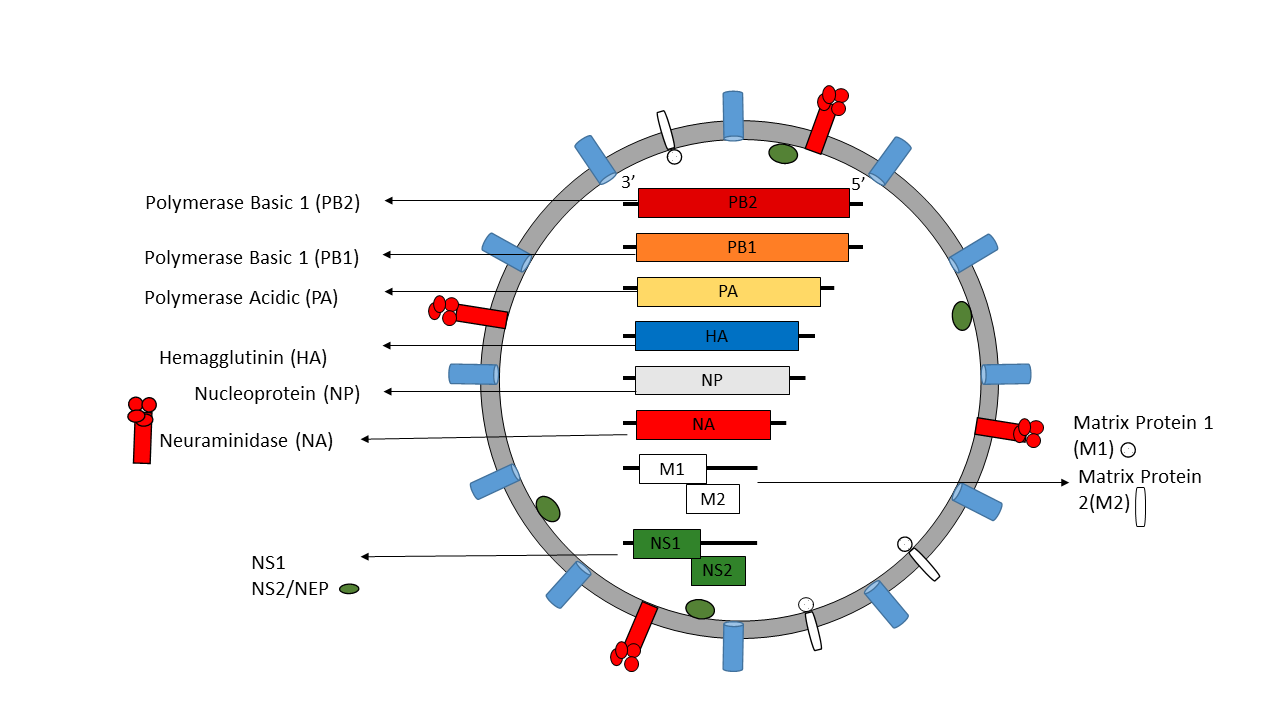
Influenza Life Cycle
IAVs initiate infection by using hemagglutinin protein (HA) to bind to cell surface glycoconjugates that contain terminal sialic acid residues, which triggers endocytosis. The resulting endosome fuses with the viral membrane and releases viral ribonucleoproteins into the cytosol, which then recruit importins that facilitate transport through the nuclear pore complex into the cell nucleus. Inside the nucleus, the viral RNA-dependent RNA polymerase then transcribes and replicates the viral RNA. The newly replicated RNA associates with nucleoprotein molecules and a single copy of the viral polymerase to assemble into a viral ribonucleoprotein, which is then trafficked to the cell surface and budded from the plasma membrane. The complex, but fast (6 hour) replication cycle involves the participation of many host proteins that are involved in multiple aspects of the viral life cycle, including viral entry, uncoating, viral RNP nuclear import, transcription and translation, post-translational modification, and transport out of the cell. The viral neuraminidase (NA) cleaves terminal sialic acids to release virions and complete the infectious cycle.
Influenza A virions possess two highly antigenic surface glycoproteins (HA and NA) that play different critical roles in the viral life cycle. Both of these proteins can be neutralized by host antibodies, and inactivated IAV vaccines induce antibody responses to HA and NA. However, because of the high mutational tolerance of these surface glycoproteins and the segmented nature of the genome that facilitates genomic reassortment during coinfection with different strains, there is opportunity for both antigenic drift and antigenic shift that allows for escape from host immune responses.
References
- WHO. Influenza. https://www.euro.who.int/en/health-topics/communicable-diseases/influenza
- Arbeitskreis Blut, Untergruppe; Influenza Virus. Transfus Med Hemother. 2009 Feb; 36(1): 32-39. doi: 10.1159/000197314
- Caini S et al. The epidemiological signature of influenza B virus and its B/Victoria and B/Yamagata lineages in the 21st century. PLoS One. 2019; 14(9):e0222381. doi:10.1371/journal.pone.0222381
- Dou, D et al. Influenza A Virus Cell Entry, Replication, Virion Assembly and Movement. Front Immunol. 2018; 9: 1581; doi: 10.3389/fimmu.2018.01581
- Glezen, WP et al. The Burden of Influenza B: A Structured Literature Review. Am J Public Health 2013; 103(3): e43-351. doi: 10.2105/AJPH.2012.301137
- Kosik, I and Yewdell, JW. Influenza Hemagglutinin and Neuraminidase: Yin-Yang Proteins Coevolving to Thwart Immunity. Viruses. 2019 Apr 16; 11(4):346. doi: 10.3390/v11040346
- Krammer, F et al. Influenza. Nature Reviews Disease Primers. 2018; 4 (3): 1-21. doi: 10.1038/s41572-018-0002-y
- Watanabe, T et al. Cellular Networks involved in the Influenza Virus Life Cycle. Cell Host & Microbe. June 2010; 7(6): 427-439. doi: 10.1016/j.chom.2010.05.008
All Orthomyxoviridae (Influenza) Products