order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Locations
Orders Processing,
Shipping & Receiving,
Warehouse
2 Shaker Rd Suites
B001/B101
Shirley, MA 01464
Production Lab
Floor 6, Suite 620
20700 44th Avenue W
Lynnwood, WA 98036
Telephone Numbers
Tel: +1 (206) 374-1102
Fax: +1 (206) 577-4565
Contact Us
Additional Contact Details
order histories, retained contact details for faster checkout, review submissions, and special promotions.
Forgot password?
order histories, retained contact details for faster checkout, review submissions, and special promotions.
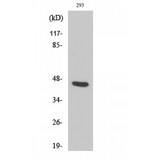
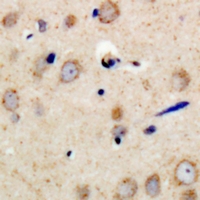
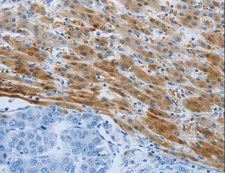
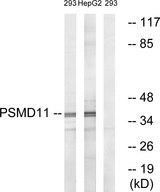
PSMD11
proteasome (prosome, macropain) 26S subunit, non-ATPase, 11
The 26S proteasome is a multicatalytic proteinase complex with a highly ordered structure composed of 2 complexes, a 20S core and a 19S regulator. The 20S core is composed of 4 rings of 28 non-identical subunits; 2 rings are composed of 7 alpha subunits and 2 rings are composed of 7 beta subunits. The 19S regulator is composed of a base, which contains 6 ATPase subunits and 2 non-ATPase subunits, and a lid, which contains up to 10 non-ATPase subunits. Proteasomes are distributed throughout eukaryotic cells at a high concentration and cleave peptides in an ATP/ubiquitin-dependent process in a non-lysosomal pathway. This gene encodes a member of the proteasome subunit S9 family that functions as a non-ATPase subunit of the 19S regulator and is phosphorylated by AMP-activated protein kinase. Alternatively spliced transcript variants have been observed for this gene.
| Gene Name: | proteasome (prosome, macropain) 26S subunit, non-ATPase, 11 |
| Family/Subfamily: | Protease Non Catalytic , not assigned-Protease Non Catalytic |
| Synonyms: | PSMD11, 26s proteasome subunit 9, S9, Rpn6, p44.5 |
| Target Sequences: | NM_002815 NP_002806.2 O00231 |
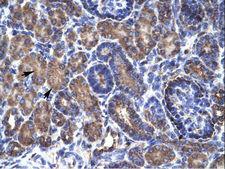
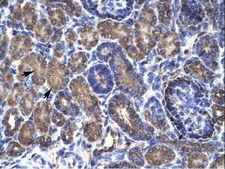
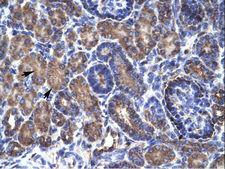
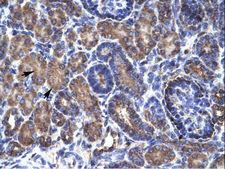
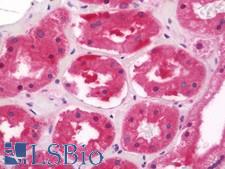
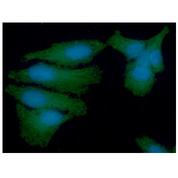
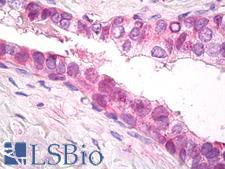
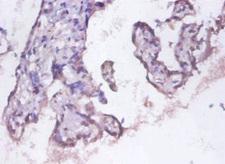

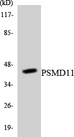


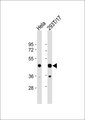
If you do not find the reagent or information you require, please contact Customer.Support@LSBio.com to inquire about additional products in development.










